![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
140 Cards in this Set
- Front
- Back
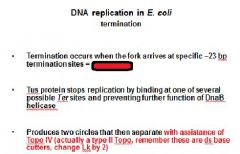

|

|
|

|

|
|
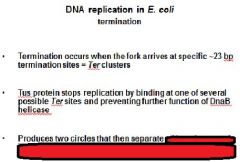

|

|
|
|
Are Ter DNA polymerase traps directional?
|
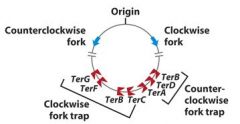
Yes
|
|
|
|
|
|
|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
Eukaryotic DNA polymerase delta is _____ slower than prokaryotic DNA pol3.
|
20x
|
|
|
|
|
|
|
|
|
|
|
|
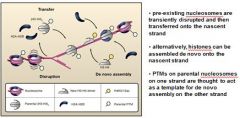
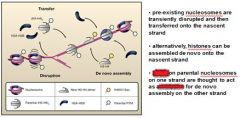
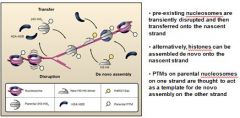
In eukaryotic DNA replication termination RNA primer degradation on the __________ strand results in a terminal gap and a ______ overhang.
|
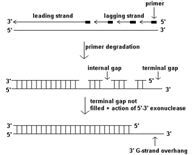
lagging, 3'
|
|
|
Each round of replication for a linear chromosome reults in?
|
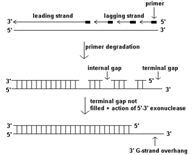
a shortening of the strands
|
|
|
The solution for dealing with the ends of linear chromosome shortening is?
|
adding a set of repeated sequences, non-encoding that can be copied using a special enzyme (telomeres)
|
|
|
What enzyme adds nucleotide bp's to a linear chromosome?
|
Telomerase
|
|
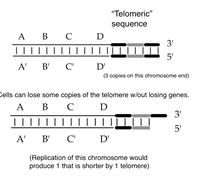
Tell me about this.
|
Yep you are right. GOOD JOB!!
|
|
|
What is the human telomere sequence?
|
TTAGGG repeated approx 1,500 times
|
|
|
In humans how many DNA lesions per person per second?
|
10,000,000,000,000
|
|
|
According to Lehninger what is the ratio of DNA lesions to mutations?
|
less than 1 in 10000
|
|
|
Ultimately what does a DNA mutation on an exon lead to?
|
alterations in a protein
|
|
|
If a basepair mutation (change) results in a change to an important amino acid for protein function is causes?
|
a deleterious mutation, often a disease state
|
|
|
A mutation is defined as a ____________ in the base sequence of ________.
|
heritable change, DNA
|
|

|

|
|

|

|
|
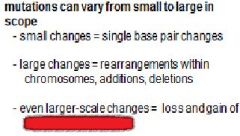
|

|
|
|
Define a silent mutation?
|
affects nonessential DNA or has a negligible effect on gene function
|
|
|
Define a nonsilent mutation?
|
They are deleterious of neutral, they are detrimental or confer no biological advantage to the affected organism
|
|
|
If a basepair change results in a change to an important amino acid for protein function it usually causes?
|
deleterious mutations and often a disease state.
|
|
|
What kind of bacterial cells are used in the Ames test?
|
mutants that are unable to produce his
|
|
|
What kind of medium is used in the Ames test?
|
His free medium
|
|
|
What is a putative mutagen?
|
putative = potential or possible
|
|
|
Describe the Ames test and how to analyze it?
|

|
|
|
The Ames test is a test for?
|
measuring of mutagenicity
|
|
|
There is a strong correlation between mutagenesis and?
|
cancer
|
|
|
Are all cancers mutagenic?
|
no
|
|
|
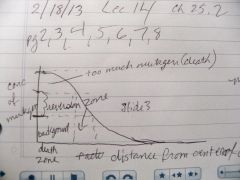
Draw a graph of the Ames test. Concentration of mutagen vs. distance from the center of disk.
Name and label the 2 zones we should know. |
|
|
|
Name the 5 repair systems we should know.
|
Methyl-directed Mismatch Repair (MMR)
Base Excision Repair (BER) Nucleotide Excision Repair (NER) Direct Repair Recombination Repair |
|
|
MMR is used to correct _______ mismatches that occur during DNA _________.
|
rare, replication
|
|
|
MMR requires what two things to function?
|
intact template (old) strand
ability to discriminate between the old and new strands (needs a hemimethylated strand? |
|
|
What is the base methylated base sequence that MMR recognizes?
|
GATC methylated a N6 of the adeninine
|
|
|
Which mut proteins form complexes that attach to mismatched DNA?
|
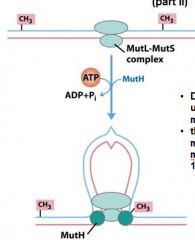
mut S
mut L |
|
|
DNA is __________ through the mutL-mutS complex until it finds ______________________________.
|
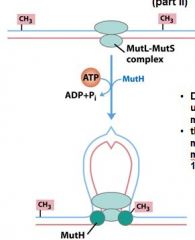
looped, methyl group on GATC
|
|
|
In the MMR repair system the distance between the mismatch and the nearest methylated adenine can be up to _________bp.
|
1000
|
|
|
Which protein in the MMR repair system acts a an endonuclease and nicks the nascent DNA at the methylated site of the parent strand?
|
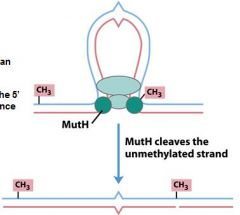
mut H
|
|
|
In MMR repair the methyl group on the parent strand activates an _______________ activity in mutH.
|
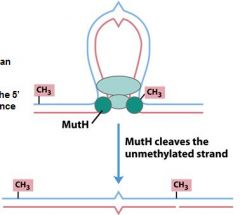
endonuclease
|
|
|
The endoncuclease activity of mutH cuts the nascent strand on the ___ end of the GATC sequence.
|
5'
|
|
|
After mutH creates a nick in the nascent strand what happens?
|
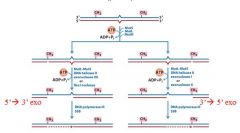
the appropriate exonuclease cuts out the bp back to the mismatch, then poly3 comes in and fixes the gap
|
|
|
Which mut protein scans the strands for a mismatch in MMR?
|
mut S
|
|
|
The MMR repair process is energetically expensive. Why?
|
up to 1000 bases are removed and replaced to correct for one mismatch
|
|
|
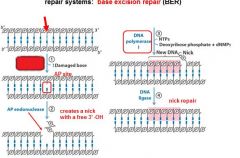
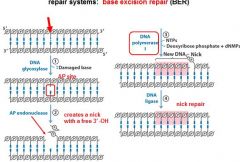
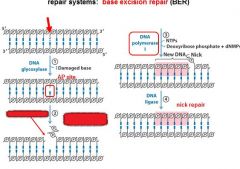
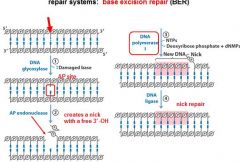
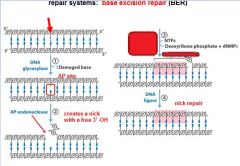
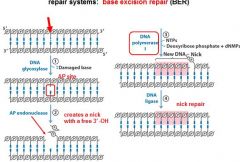
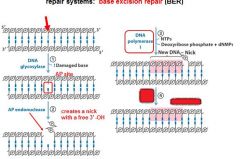
The BER replaces base lesion due to?
|

|
|

|

|
|

|

|
|

|

|
|

|

|
|

|

|
|
|
DNA glycoslyases in the BER system hydrolyze the _____________ bond.
|
N-glycosidic bond
|
|
|
DNA glycoslyases in the BER system are ____________ for a given base.
|
specific
|
|
|
DNA glycoslyases in the BER system are specific for _________ and ____________.
|
a given base, also specific to damaged type of bases
|
|
|
|
|
|
|
|
|
|
|
|
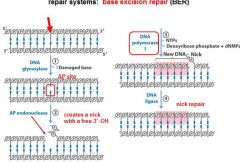
|
|
|
What does MMR, BER, and NER stand for in DNA repair systems?
|
methyl-directed mismatch repair
base excision repair nucleotide excision repair |
|
|
NER is used to repair ___________ lesions that cause _____________ changes in DNA.
|
bulky, conformational
|
|
|
What are two kinds of bulky lesions?
|
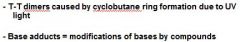
|
|
|
Name two causes of base adducts.
|
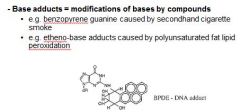
|
|
|
What is unique about the nuclease activity of NER?
|

|
|
|
Mutations in the exinuclease of the homologous human pathway of NER can cause?
|

|
|
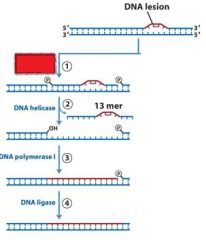
|
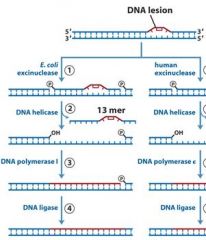
|
|
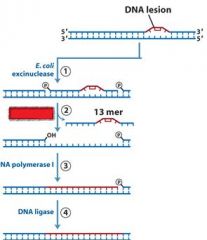
|
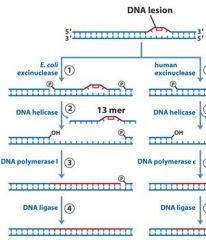
|
|
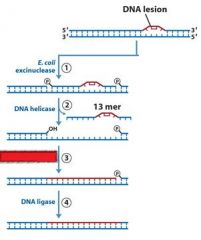
|
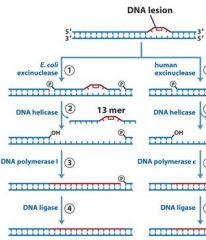
|
|
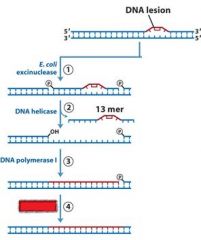
|
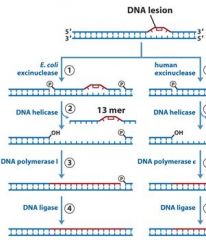
|
|
|
MMR uses DNA poly _____, while NER and BER uses DNA poly ____.
|
3,1
|
|
|
In the BER repair system a damaged base is recognized by a ______________________.
|
DNA glycosylase
|
|
|
Direct nucleotide repair involves direct repair of a _________ base without ___________ if the phosphodiester backbone.
|
defective, cleavage
|
|
|
Direct nucleotide repair involves direct repair of a defective base without cleavage of the _____________ ____________.
|
phosphodiester backbone
|
|

Some examples of direct nucleotide repair.
|

|
|

Some examples of direct nucleotide repair.
|

|
|

Some examples of direct nucleotide repair.
|

|
|

Some examples of direct nucleotide repair.
|

|
|

Some examples of direct nucleotide repair.
|

|
|

Some examples of direct nucleotide repair.
|

|
|
|
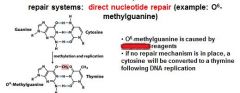
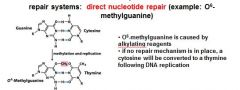
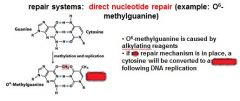
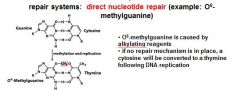
In the event of an O6-methylated guanine the first round of replication will produces?
|
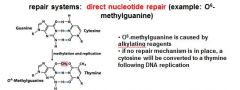
G-T
|
|
|
|
|
|
|
|
|
O6-methylguanine is corrected in the direct nucleotide repair system by_____________________________.
|

|
|
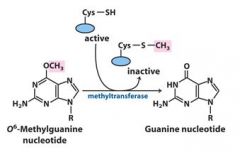
This repair mechanism is in which repair system? This reaction is not __________, and not ___________.
|
direct nucleotide repair, catalytic, reversible
|
|
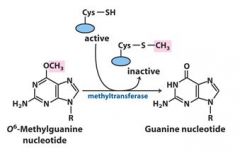
This direct nucleotide repair mechanism is considered energetically expensive because?
|

|
|
|
The recombination repair method is different from MMR, NER, and BER because?
|
it does not require a template
|
|
|
If an O6-methylguanine is left uncorrected in DNA it will convert a ____ pair to a _____ pair.
|
G-C to A-T
|
|
|
Are double strand breaks (DBS) common in DNA?
|
yes
|
|
|
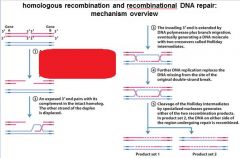
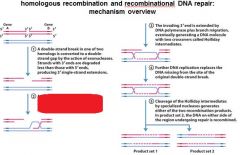
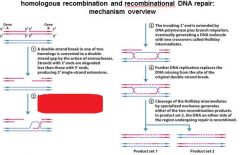
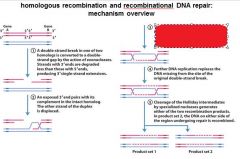
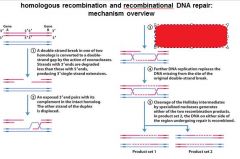
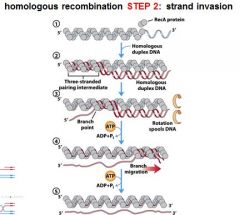
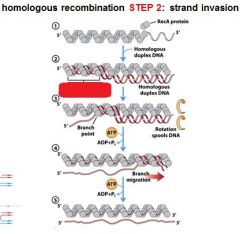
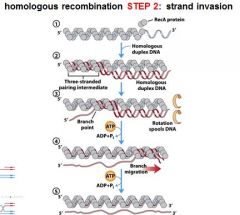
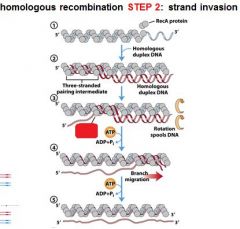
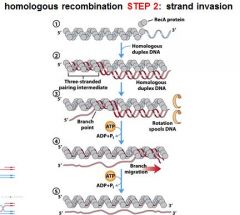
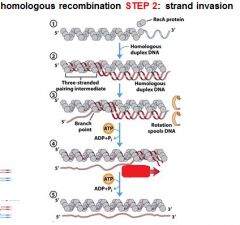
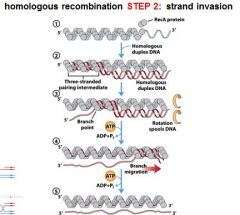
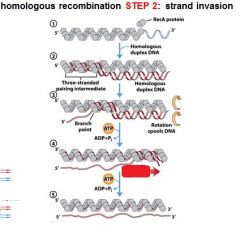
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the __________ into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
DSB (double strand break)
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the ___________ complex 2. Strand invasion via the recA protein complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
recBCD
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. via the recA protein complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
Strand invasion
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the ____________ complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
recA protein
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. ____________; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
3' end extension
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. 3' end extension; branch migration 4. ______________ 5. resolution of Holliday intermediate by RuvC |
3' end extension
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of __________ intermediate by RuvC |
Holliday
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. 3' end extension; branch migration 4. 3' end extension 5. resolution of Holliday intermediate by ________ |
ruvC
|
|
|
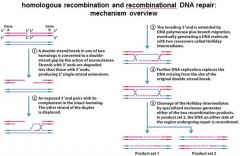
|
|
|
|
|
|
|
|
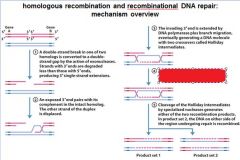
|
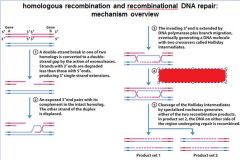
|
|
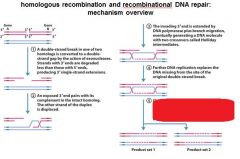
|
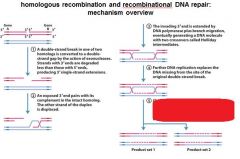
|
|

This is called?
|
D-loop
|
|

This is called?
|
double holliday junction or intermediate
|
|
|
5 STEPS IN DNA RECOMBINATION REPAIR:
1. Expanding the DSB (double strand break) into a gap by the RecBCD complex 2. Strand invasion via the recA protein complex 3. 3' end extension; ___________________ 4. 3' end extension 5. resolution of Holliday intermediate by RuvC |
branch migration
|
|
|
In DNA recombination the _____________ has both helicase and nuclease activities.
|
RecBCD complex
|
|
|
In DNA recombination the RecBCD complex has both __________ and _________ activities.
|
helicase, nuclease
|
|
|
In DNA recombination the RecBCD complex initiates DSB repair by attaching to and degrading free ________________.
|
double strand ends
|
|
|
In DNA recombination the ________ complex initiates DSB repair by attaching to and degrading free double strand ends.
|
RecBCD
|
|
|
In DNA recombination the RecBCD complex initiates DSB repair by ________ to and _________ free double strand ends.
|
attaching, degrading
|
|
|
What is the purpose of the chi-scanning site in the RecBCD complex?
|
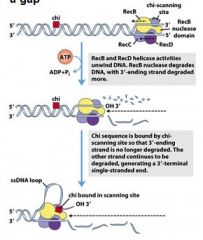
enzyme binding to the chi sequence halts degradation of the 3'end while degradation of the 5'end continues
|
|
|
In DNA recombination repair what step is the RecBCD complex involved in?
|
1. expanding the DSB into a gap
|
|
|
Does more bp per turn mean overwinding or underwinding?
|
underwinding
|
|
|
The RecA protein complex is involved in what step of DNA recombination repair?
|
2. strand invasion
|
|
|
The RecA protein polymer form ______________ helical polymer filaments that fit into the ________ groove of DNA molecules.
|
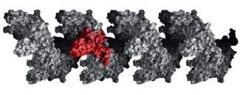
right-handed, major
|
|
|
The RecA protein polymer assembles on an existing ________ in the 5'--3' direction.
|
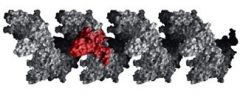
ssDNA
|
|
|
The RecA protein polymer assembles on an existing ssDNA in the _______ direction.
|
5'--3'
|
|
|
RecA can facilitate ______________ and _____________.
|
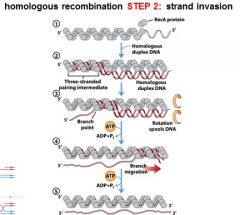
strand invasion, branch migration
|
|
|
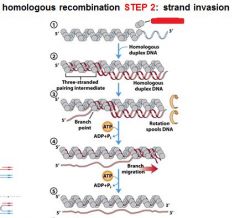
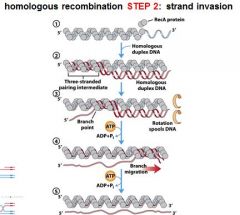
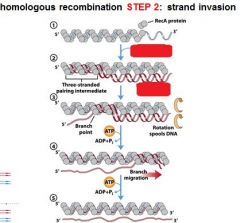
The RecA protein polymer can mediate strand invasion only if presented with _________.
|
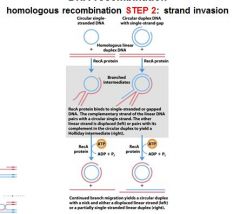
ssDNA
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
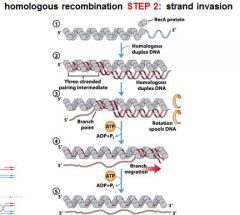
|
|
|
In DNA recombination repair what protein complex is associated with step 1: expanding the DSB into a gap.
|
RecBCD
|
|
|
In DNA recombination repair what protein complex is associated with step 2: strand invasion?
|
RecA
|
|
|
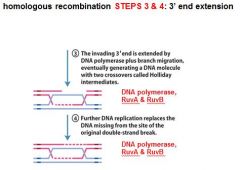
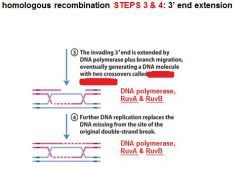
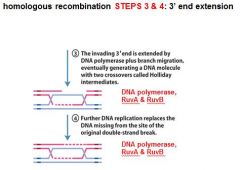
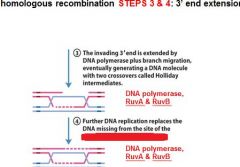
In DNA recombination repair what enzymes are associated with step 3: 3' end extension branch migration?
|
DNA polymerase, RuvA, RuvB
|
|
|
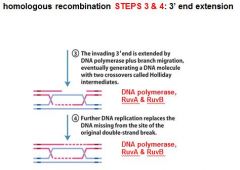
In DNA recombination repair DNA polymerase, RuvA and RuvB are associated with what step(s)?
|
3. 3' end extension: branch migration
4. 3' end extension |
|
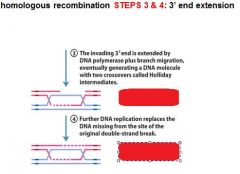
Enzymes?
|
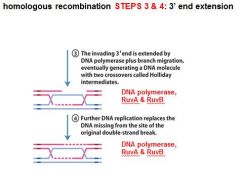
|
|
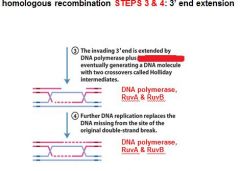
|
|
|
|
|
|
|
|
|
|
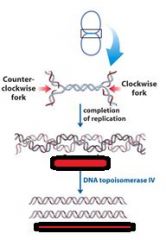
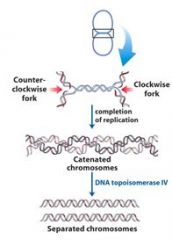
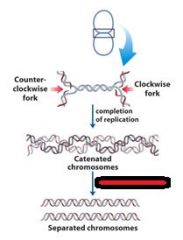
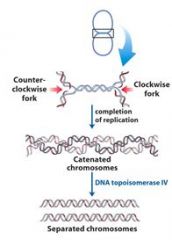
Define concatenation?
|
replication of a circular chromosome results in two topologically interlinked (cantenated) circular chromosomes. DNA circles linked in this way are known as CATENANES.
|

