![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
74 Cards in this Set
- Front
- Back
|
Nuclear envelope |
Inner and outer membrane Continuous with ER |
|
|
Nuclear lamina |
Sheet like network of intermediate filaments that line the inner membrane of nucleus Provide structural and provide sites of chromatin attachment |
|
|
Nuclear pore complex |
Highly complex proteinaceous pore that regulates the entry/exit of proteins and exits of mRNA |
|
|
Nucleolus |
Ribosome-producing sub-compartment of the nucleus |
|
|
Nucleoplasm |
Chromatin/chromosomes containing region |
|
|
Chromatin |
Chromosomal material- is a protein-dna complex (plus some RNA) |
|
|
Histones function |
Package and order DNA into structured units |
|
|
When are histones made? |
G1 phase |
|
|
S phase |
DNA is replicated and histones and non histones proteins are deposited on the daughter dna molecules to produce chromatin structures |
|
|
G2 phase |
DNA content is doubled |
|
|
Mitosis |
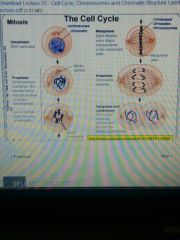
|
|
|
What does chromatin consist of? |
Dna, protein and small amount of rna |
|
|
Nucleosomes |
Histones package and order of DNA into structured units Beads on a string |
|
|
5 types of histones |
H2a H2b H3 H4 Core histones - 2 copies of each form an octamer (8 subunits) H1 Clamps DNA wrapped around nucleosomes |
|
|
Which type of histones is highly conserved? |
H3, H4 |
|
|
What does histones contain? |
Small, basic proteins - contains 25% lys and arg Why? Neutralize highly charged dna |
|
|
Nucleoplasmin |
Molecular chaperone Assembly of histones with dna |
|
|
Nucleosomes are deposited in stepwise manner |
1) tetramer of two h3 and 2 h4 dimer are recruited first 2) followed by h2a-h2b dimer 3) h1 clamps DNA wrapped around nucleosomes (~1.65 loops) |
|
|
What digest the linker DNA? |
Nuclease |
|
|
Linker DNA |
Beads on a string form of chromatin |
|
|
What is the average length of linker DNA? |
~ 53 nt-pairs |
|
|
Diameter of nucleosomes |
11nm |
|
|
How to separate histones and DNA? |
High concentration of salt |
|
|
What is length of DNA after histones is taken off? |
147 nt pairs DNA double helix |
|
|
Lk is a _______ property of circular dna |
Topological Doesn't change when dna is bent or deformed unless one or both strands is broken |
|
|
Left handed, solenoidal supercoil |
DNA wraps around the nucleosomes and changes lk by -1 |
|
|
Left handed plectonemic supercoil |
Unbound positive supercoil and linking number (+1) |
|
|
What can relax the positive plectonemic supercoil causing a net negative supercoil? |
Topoisomerase 1 |
|
|
In nucleosomes, what type of supercoil wraps around histones octamer? |
Left handed, solenoidal supercoil |
|
|
What is the 2nd level of chromatin organization? |
30nm fiber or solenoids 11nm chromatin fibers coil to form 30 nm fiber |
|
|
Wrapping the DNA around histones core compact the DNA ______ |
~7 folds |
|
|
Where h1 located on the 30nm fiber ? |
Interior |
|
|
Does compaction occur throughout entire chromosome ? |
No. Regions in which gene are being actively transcribed are less orders and have little to no h1 |
|
|
What does compaction make DNA inaccessible to ? |
Transcription and replication factors |
|
|
Where on DNA contains little h1? |
Regions that are Transcriptionally active- less order |
|
|
Does transcription occur in solenoid? |
No |
|
|
Scaffold |
Associated regions are separated by loops of dna |
|
|
Higher order chromatin |
Coils upon coils Loops of DNA may contain related genes |
|
|
What are the three levels of DNA packing? |
1) nucleosomes 2) 30nm fiber 3) extended form of chromosome Condensed section of chromosomes Mitotic chromosomes |
|
|
Where does histones interact primarily with? |
Sugar phosphate backbone Interaction not sequence specific |
|
|
Why cant ~70 % DNA surface exposed and available to interact with DNA binding protein? |
Inaccessible due to dna/nucleosomes binding |
|
|
What does n-terminal tails protrude from? |
Octamers |
|
|
Structural organization of histones |

|
|
|
Handshake interaction |
Histone 2a and 2b form a dimer |
|
|
What interaction does a h3 and h4 form? |
Dimer All together to form tetramer |
|
|
H3/h4 or h2a/h2b forms a scafford? Forms a octamer? |
H3/h4 h2a/h2b onto H3/h4 |
|
|
Why are n terminals important? |
Assembly for 30 nm fiber and gene regulation |
|
|
Why in xray crystal structure are most histones n terminals not visible? |
Suggests they are flexible |
|
|
Properties of n terminals tails of histones (4) |
Protrude from octameric disk Highly conserved between histones Positively charged - lys and are Highly flexible |
|
|
What does n terminal histones tails mediate? |
30 nm fiber assembly |
|
|
How do histones tails help pack nucleosomes into 30 nm fiber? |
+ charged n termini bind - sugar-phosphate backbone on DNA of neighboring nucleosomes |
|
|
Chromatin remodeling |
Increase or decrease its accessibility to proteins in cell Can change nucleosomes structure and modify histones |
|
|
What does chromatin remodeling complexes require? |
Atp |
|
|
Exposed DNA can accessed by enzymes involved in? (4) |
Gene expression, DNA replication , recombination and repair |
|
|
What n terminal histones tails are subject to covalent modification? |
Reversible acetylation of lysines (-coch3), phosphorylation of serines (-po4), methylation of lysines (-ch3) |
|
|
Acetylation of lysines |
Acetylation core histones (h3/h4) Neutralizes positive charge of lysine residues which reduce binding to DNA and destabilizes chromatin Likely facilitates the of extended regions consisting of the 11nm fiber
|
|
|
Methylation of lysines |
Three different level of lysine methylation can occur - all results in lysine positively charged Depending on which residue is modified, methylation can be associated with gene activation or gene silencing |
|
|
Serine phosphorylation |
Add a neg charge to histones Impact of histones phosphorylation in terms of chromatin stricture/gene expression is somewhat less well defined |
|
|
Histone code |

|
|
|
Heterochromatin |
Tightly packed form of DNA Stains darker Some regions of DNA are consistently compacted as heterochromatin (constitutive heterochromatin) Commonly found near centrosomes and telomeres |
|
|
Euchromatin |
Lightly packed form of DNA Stains lighter Often under active transcription Predominant form during interphase |
|
|
What happens in chromosome remodeling during DNA replication ? |
As replication fork approaches a stretch of chromatin, nucleosomes on parental duplex is displaced with new ones on reformed on newly synthesized daughter duplexes |
|
|
What 3 kinds of DNA sequences within the chromosomes is controlled for replication? |
Replication origin Centromere Telomere |
|
|
Kinteochore |
Serves as attachment site for mitotic spindle (protiens complex) |
|
|
Centromere |
Part of a chromosome that link two sister chromatid Allows one copy of each duplicated and condensed chromosome to be pulled to each daughter cell during cell division. |
|
|
Operon |
Genes that encode protiens that are involved in the same process |
|
|
Structural genes |
Gene that codes for any RNA or protiens product other than a regulatory factor |
|
|
What does a operon encompass? |
Promoter, operator and structural genes |
|
|
Polycistronic mrna |
Genes are transcribed together onto a single mRNA |
|
|
Cistron |
Each section of these mRNA may then be translated independently |
|
|
Effector molecules |
Transcription of operon can be induced or repressed |
|
|
Negative regulation |
Repressor protiens binds to DNA at a site or near promoter and blocks transcription by RNA pol |
|
|
Positive regulation |
An activator protein binds to DNA at activator binding sites and enhances the activity of RNA pol |
|
|
Where are repressors and activator protiens bound for turning promoter on and off? |
Operator and activator binding sites |

