![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
78 Cards in this Set
- Front
- Back
|
Do Microbes Make Unnecessary Proteins? |
No, the microbe will not waste energy making unneeded proteins. - It uses elegant mechanisms to control gene expression at various levels. |
|
|
Central Dogma Visual |
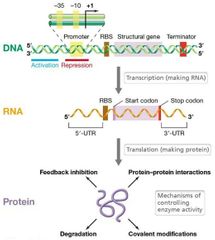
|
|
|
The two Approaches to Regulation |
Regulation of gene expression... - transcription initiation. - transcription elongation. - translation. & Alter activity of enzymes and proteins... - posttranslational. |
|
|
The 2 different outcomes from DNA binding... |
The binding event can block transcription (negative regulation).
The binding event can activate transcription (positive regulation). |
|
|
Def: Negative Control |
Negative control: a regulatory mechanism that stops transcription. |
|
|
Def: Repression |
Preventing the synthesis of an enzyme in response to sufficient amounts of a product. - specific effect. - widespread control for amino acid and nucleotide precursors. - Usually, final product of a biosynthetic pathway represses enzymes. - Usually affects biosynthetic anabolic enzymes. |
|
|
Repression Visual |
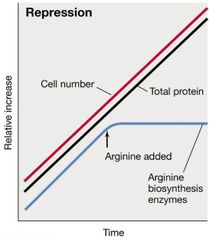
|
|
|
Negative Control: Induction |
The production of an enzyme in response to presence of substrate. - Typically affects catabolic enzymes (e.g., lac operon). - Ensures enzymes are synthesized only when needed. |
|
|
Induction Visual |
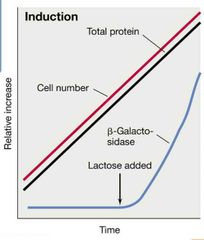
|
|
|
Oppressions: Repression Visual |
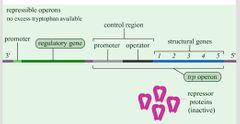
|
|
|
Oppressions: Induction Visual |
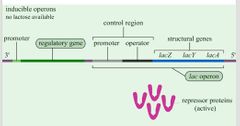
|
|
|
Negative Control: Inducer |
Substance that induces enzyme synthesis.
|
|
|
Negative Control: Corepressor |
A substance that represses enzyme synthesis. |
|
|
Negative Control: Effectors |
Collective term for inducers and corepressors. (Basically anything that "effects" an enzyme) - typically small molecules. - can be structural analogs of substrates/products, (e.g., allolactose). |
|
|
Effectors and Transcription |
Effectors indirectly affect transcription by binding to specific DNA-binding proteins. - Corepressors bind to an allosteric repressor protein. - Allosteric repressor is activated and binds to region of DNA near promoter called the operator. |
|
|
Corepressor and Repressor Visual |
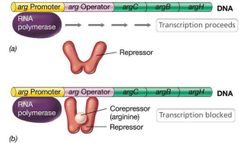
|
|
|
Def: Operon |
Cluster of consecutive genes whose expression is under control of a single operator. - All genes transcribed as single mRNA. - Transcription physically blocked when repressor binds to operator. |
|
|
Enzyme Induction |
Enzyme induction can also be controlled by a repressor. - Addition of inducer inactivates repressor, and transcription can proceed. Repressor's role is inhibitory (preventing mRNA synthesis), so it is called negative control. |
|
|
Repressor and Inducer Visual |
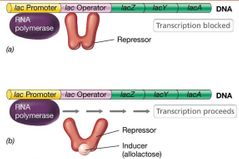
|
|
|
Def: Positive Control |
Regulator protein that activates the binding of RNA polymerase to DNA.
DNA Activator proteins bind specifically to activator-binding site (certain DNA sequence that is not called an operator). |
|
|
Ex of Positive Control: Maltose & E. Coli |
Maltose catabolism in E. Coli - Maltose activator protein cannot bind to DNA unless it first binds maltose (inducer). - Subsequent binding. |
|
|
Maltose and Inducer Visual |
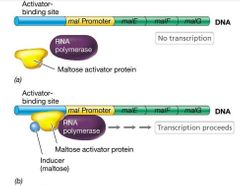
|
|
|
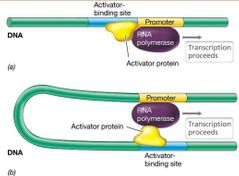
Activation of Positive Control |
Promoters of positively controlled operons only weakly bind RNA polymerase.
Activator protein helps RNA polymerase recognize promoter. - May bend DNA structure - May interact directly with RNA polymerase.
Many operons have multiple types of control. |
|
|
Activator Protein & Positive Control Visual |
|
|
|
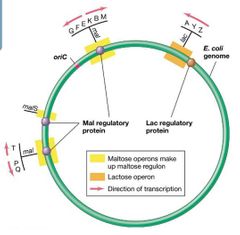
Regulons |
Genes for maltose are spread out over the chromosome in several operons. - Each operon has an activator-binding site. - Multiple operons controlled by the same regulatory protein are called a regulon.
Regulons also exist for negatively controlled systems (e.g., arginine regulon). |
|
|
Operon Visual |
|
|
|
Key Components of Operons: Promoter |
Site on the DNA bound by the RNA polymerase; directs the initiation of transcription. |
|
|
Key Components of Operons: Activator |
Protein that binds to a site on the DNA; assists binding of the RNA polymerase to the promoter, resulting in increased transcription initiation. |
|
|
Key Components of Operons: Activator Binding Site |
Site on the DNA bound by the activator. |
|
|
Key Components of Operons: Repressor |
Protein that binds to the operator site on the DNA, inhibiting transcription. |
|
|
Key Components of Operons: Operator |
Site on the DNA bound by the repressor. |
|
|
Key Components of Operons: Effector |
Small molecule thst binds to activator or repressor proteins, modifying their gene regulation activity. |
|
|
Key Components of Operons: Inducer |
Effector that increases transcription by either enabling an activator or disabling a repressor. |
|
|
Key Components of Operons: Corepressor |
Effector that decreases transcription by enabling a repressor. |
|
|
Def: Global Control Systems |
Regulate expression of many different genes simultaneously (e.g., lactose operon and maltose regulon). |
|
|
Ex of Global Control System |
Catabolite repression is an example of global control. - Controls use of carbon sources if more than one present. - Synthesis of unrelated catabolic enzymes (e.g., lactose operon and maltose regulon) is repressed if glucose is present in growth medium. - Also called “glucose effect”. - Ensures that the "best" carbon and energy source is used first. |
|
|
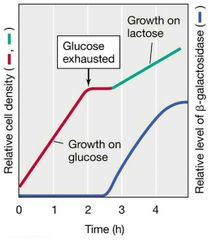
Diauxic Growth |
Two exponential growth phases if two energy sources available. - Better energy source consumed first, growth stops. - After lag, growth resumes with second energy source. (When controlling the aux cord, sometimes you have a lull inbetween playing two dope songs. The first one was clearly the best though). |
|
|
Diauxic Growth Visual |
|
|
|
CRP |
CRP = Cyclic AMP Receptor Protein. In catabolite repression, transcription is controlled by the (CRP), an activator protein, a form of positive control.
CRP binds to DNA only if it has bound cyclic adenosine monophosphate (cyclic AMP or cAMP), a regulatory nucleotide derived from a nucleic acid precursor (ATP) |
|
|
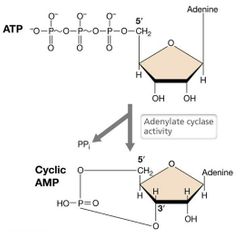
Cyclic AMP |
Cyclic Adenosine Monophosphate - Cyclic AMP or cAMP. Regulatory nucleotide derived from a nucleic acid precursor (ATP). Permits the binding of CRP to DNA. It must be bound first. |
|
|
Cyclic AMP Structure Visual |
|
|
|
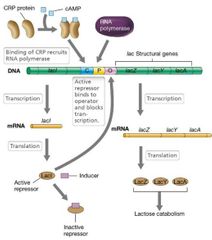
For lac genes to be transcribed... |
Cyclic AMP level must be high enough for CRP protein to bind to CRP-binding site.
Lactose or another inducer must be present to prevent lactose repressor (LacI) binding. |
|
|
Cyclic AMP Pathway Visual |
|
|
|
Prokaryotes and their Environments |
Prokaryotes regulate cellular metabolism in response to environmental fluctuations. - External signal may be transmitted directly to the target. - External signal may be detected by sensor and transmitted to regulatory machinery (signal transduction). • Most signal transduction systems are two-component regulatory systems. |
|
|
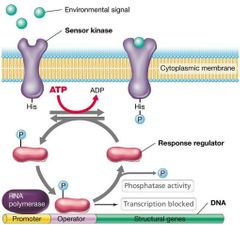
Two-Component Regulatory Systems |
Made up of two different proteins - Sensor kinase (in cytoplasmic membrane): detects environmental signal and autophosphorylates. - Response regulator (in cytoplasm): DNA-binding protein that regulates transcription. |
|
|
Feedback Loop |
Terminates signal. Uses phosphatase that removes phosphate from response regulator. |
|
|
Feedback Visual |
|
|
|
Chemotaxis |
The ability of an organism to sense chemical gradients and modify its motility in response. - It is a behavior in which motile bacteria swim toward favorable environments (chemoattractants) or away from unfavorable environments (chemorepellents). |
|
|
Modified Two-Component System |
Used in chemotaxis to... - Sense temporal changes in attractants or repellents. - Regulate flagellar rotation. - Thus regulate activity of preexisting proteins instead of modifying transcription of genes. |
|
|
Def: Sensory Proteins |
Located in the cytoplasmic membrane, they sense attractants and repellents, and interact with cytoplasmic sensor kinases. |
|
|
Def: MCPs |
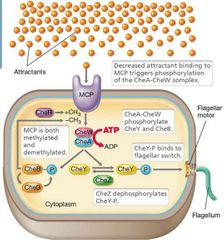
Methyl-accepting chemotaxis proteins (MCPs) - Bind attractant or repellent and initiate flagellar rotation. - Interact with CheA (sensor kinase) and CheW. |
|
|
Def: CheA |
The cytoplasmic domains of each MCP binds to the protein CheA and controls its activity.
CheA works as the sensor kinase, becoming phosphorylated.
CheA then phosphorylates CheY (the Response Regulator or RR protein).
- Phosphorylated RR proteins do not bind DNA; they bind to the flagellar motor, changing its activity. |
|
|
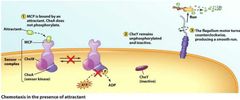
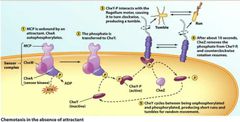
The direction of Flagellar Motor Rotation... |
...determines the type of movement: Controlled by CheY protein. - CheY = counterclockwise and smooth swimming). - CheY-P (phosphorylated) = clockwise and random. Forms when no atractants are present in environment. - Random movement = drop in CheY-P, CheY returns along with CCW and smooth movement. - CheZ dephosphorylates CheY-P. |
|
|
Chemotaxis w/ Attractant Visual |
|
|
|
Chemotaxis w/out Attractants Visual |
|
|
|
Fx. of Chemotaxis Proteins Visual |
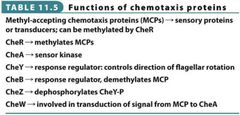

|
|
|
CheR |
Methylates MCPs. |
|
|
CheA |
Sensor Kinase. |
|
|
CheY |
Response regulator: controls direction of flagellar rotation. |
|
|
CheB |
Response regulators, demethylates MCP. |
|
|
CheZ |
Dephosphorylates CheY-P |
|
|
CheW |
Involved in transduction of signal from MCP to CheA. |
|
|
Chemotaxis Adaptation |
Stop responding and reset. |
|
|
Chemotaxis Feedback Loop |
Allows the system to reset itself to continue to sense the presence of a signal.
- Relies on response regulator CheB.
- Involves modification of MCPs: methylation stops response to attractants and increases response to repellants.
- Reversible methylation or demethylation of MCPs desensitizes or sensitizes MCPs, respectively. |
|
|
Chemotaxis Visual |
|
|
|
Do Prokaryotes communicate with one another? |
Prokaryotes can respond to the presence of other cells of the same species. |
|
|
Quorum Sensing |
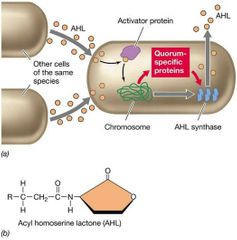
Mechanism by which Bacteria and some Archaea assess their population density. Ensures that a sufficient number of cells are present before initiating a response that, to be effective, requires a certain cell density (e.g., toxin production by pathogenic bacterium). |
|
|
Autoinducer |
A specific signaling molecule that each species of bacterium produces. Diffuses freely across the cell envelope. Reaches high concentrations inside cell only if many cells are nearby and making the same autoinducer. Binds to specific activator protein or sensor kinase, triggering transcription of specific genes. |
|
|
Quorum Sensing Visual |
|
|
|
Virulence Factors |
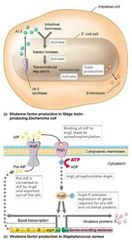
Ex: Escherichia coli O157:H7 - Shiga toxin–producing strain. - Produces AHL AI-3 that induces virulence genes. - Epinephrine plus norepinephrine plus AI-3 bind to sensor molecules in plasma membrane. • Activates motility, toxin secretion, and production of lesion-forming proteins. |
|
|
Virulence Factors Visual |
|
|
|
Stringent Response |
Used to survive nutrient deprivation, environmental stress, and antibiotics. Shuts down macromolecule synthesis and activates stress survival pathways. |
|
|
Stringent Response - E. Coli |
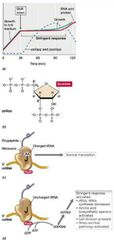
AA excess -> limitation = rRNA + tRNA + ribosome synthesis stops.
Protein + DNA synthesis stop, new AAs biosynthesized.
Later, rRNA + ribosome prod. begins but at much lower rate.
Triggered by (p)ppGpp: the alarmones guanosine tetraphosphate (ppGpp) and guanosine pentaphosphate (pppGpp).
RelA and SpoT proteins also important. |
|
|
Stringent Response Visual |
|
|
|
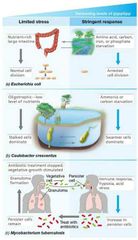
Stringent Response and Microbial Ecology (Part 1) |
Environment/habitat determines response. Ex: voiding E. coli in feces reduces nutrients, initiates ppGpp synthesis, stringent response occurs. Ex: Caulobacter stringent response triggered by carbon/ammonia starvation instead of amino acid limitation. - ppGpp increases swarmer (motile) cell formation, may reach a niche with more nutrients. |
|
|
Stringent Response and Microbial Ecology (Part 2) |
Ex: Mycobacterium tuberculosis lungs hypoxic and phosphate-limited, triggering stringent response. Converts a subpopulation of dormant persister cells resistant to antibiotics that can revert back to infective cells. |
|
|
Limited vs. Stringent Response Visual |
|
|
|
|

