![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
20 Cards in this Set
- Front
- Back
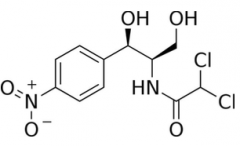
Chloramphenicol |
Structure: D-Threo (NH-Cl2) is the active form. Look for Cl2 Ribosome MOA: Binds to the 50S portion and inhibits peptide bond formation Resistance: Production of chloramphenicol acetyltransferase (CAT), which acetylates chloramphenicol making it unable to bind to the ribosome. |
|
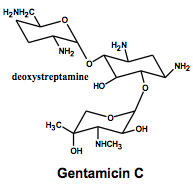
Aminoglycosides Kanamycin B, Gentamicin C, Amikacin, Tobramycin |
Ribosome Structure: Amino sugars linked glycosidically. Stable at pH2-11 MOA: Binds to the 30S subunit, causing misreading of genetic code Do not easily cross cell wall so they are given in combination with a cel wall biosynthesis inhibitor (ex. penicillin) Resistance: Enzymes that adenylate or phosphorylate hydroxy groups or Enzymes that acetylate amino groups |
|
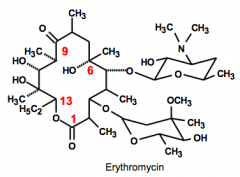
Macrolides Erythromycin |
Ribosome Structure: 12- 14 Lactone (cyclic ester) ring. Unstable basic compound MOA: Binds to the 50S subunit and prevents translocation Resistance: Specific methylation of an evening residue at the erythromycin-binding site on ribosome. |
|
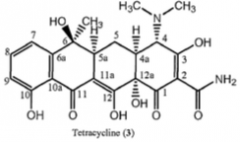
Tetracycline Chlorotetracycline, Oxytetracycline |
Ribosome Structure: 4 rings MOA: Binding to 30S subunit and prevent binding of aminoacyl t-RNA. Resistance: Modification of ribosome structure or Synthesis of new bacterial enzymes that cause pumping out of the antibiotic |
|
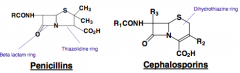
Beta-Lactams |
Cell Wall Structure: 4-membered amide (lactam fused to 5- membered (penicillins) or 6-membered (cephalosporins) dihydrothiazine ring. MOA: Mimics D-Ala-D-Ala and binds to and inactivates transpeptidase (bind to active site of PBPs) involved in the final cross-linking of peptide bridges with neighboring monomers. Resistance: Alterations in the penicillin binding proteins structure or Secretion of beta-lactamase enzyme which breaks open the beta lactic ring rendering the molecule ineffective in binding to the transpeptidases Combat: Usually administered in combination with clavulamic acid or Administered with structural modifications that make them more resistant to the action of beta-lactamases. |
|
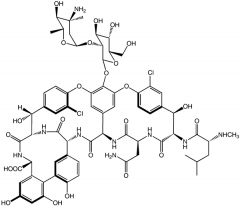
Vancomyocin |
Cell wall Structure: Active on Gram-postivie bacteria MOA: Ties up D-Ala-D-Ala so that it cannot interact with transpeptidase. Resistance: Bacteria synthesizes D-Ala-D-Lac which no longer binds to vancomyosin. |
|
|
Isoniazid (INH) |
Cell Wall Structure: Activated by the oxidative enzyme KatG to a radical form, which reacts with NAD+. MOA: Converted to reactive nitrogen species which inhibits formation of mycelia acid, a key component of the cell walls of mycobacterium.
|
|
|
Daptomycin |
Cell Wall Structure: Cyclic anionic lipopeptide isolated from streptomycin roseosporus. Consists of 13 AA residues. MOA: In the presence of Ca2+ it binds to glycero-3-phosphocholine and experiences a conformational change which increases amphipathicity, decreases charge, and allows it interact with neutral or acidic membranes forming micelles. then In the presence of Ca2+ major conformational change allows daptomycin to insert into bilayer membrane with acidic character this leads to depolarization and loss of K+ ions and cell death |
|
|
Thiocaramate and Allylamine |
Antifungal Supression of fungal squalene epoxidase to result in accumulation of squalene and decreased ergosterol |
|
|
Benzofuran cyclohexene |
Antifungal Binding to fungal DNA to cause malformation of spindle and cytoplasmic microtubules |
|
|
Polyene |
Antifungal Binding to ergosterol in fungal cell membranes to result in membrane disorganization |
|
|
Pyrimidine |
Antifungal Deamination by fungal cells to 5-flurouracil which is incorporated into RNA in place of uracil or is converted to 5-flurouracil-2'-deoxyuridylic acid which inhibits thymidine synthetase |
|
|
Azole |
Antifungal Inhibition of cytochrome P-450 that catalyzes 14a-demethylation of lanosterol to ergosterol accumulation of 14-methylated sterols cause permeability disruption |
|
|
Relenza and Tamiflu |
Anti-viral Structure: Plug Drugs MOA: Nuraminidase Inhibitors, mimic sialic acid (carbohydrate) and bind strongly to the sialidase enzyme preventing its function (breaks down unwanted sialic acid) |
|
|
Amantadine and Rimantadine |
Antiviral Inhibit viral entry into cell and also uncoating of viral sheath proteins Against influenza type A |
|
|
Acyclovir (HSV, and VSV) and Ganciclovir (also CMV) |
Antiviral Nucleoside antimetabolites (inhibitor of DNA polymerase) Herpes simplex virus and Varicella Zoster virus Virus specified enzyme converts acyclovir into eh monophosphate and then host kinases convert further in di- and tri-phosphates |
|
|
Vidarabine |
Antiviral Nucleoside antimetabolites (inhibitor of DNA polymerase)Developed as a potential anti-cancer agent initially Arabinose sugar in place of the deoxyribose and this slow down viral DNA polymerase |
|
|
Metronidazole No MOA |
Protozoa and Bacteria Reductive bioactivation of its nitro goroup to form reactive cytotoxic products that interfere with nucleic acid synthesis |
|
|
Mupirocin No MOA |
Gram + cocci inhibits protein synthesis by selectively binding to isoleucyl-tRNA synthetase. |
|
|
Polymyxins No MOA |
Bactericidal against gram - bacteria Polypeptides Cationic detergents, disrupting bacterial cell membranes Bind and inactivate endotoxin |

