![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
62 Cards in this Set
- Front
- Back
|
Fate mapping
|
fate mapping is a method of understanding the embryonic origin of various tissues in the adult organism by establishing the correspondence between individual cells (or groups of cells) at one stage of development, and their progeny at later stages of development.
|
|
|
Fertilization
|
egg and sperm meet to form zygote |
|
|
Cleavage
|
fast divisions, no growth |
|
|
Gastrulation
|
folding of tissue, formation of germ layers |
|
|
Organogenesis |
creation of organs from germ layers
|
|
|
Metamorphosis
|
Period of growth and development after larval stage before reaching maturity
|
|
|
Gametogenesis
|
formation and maturation of germ cells
|
|
|
embryotic stages
|
fertilization cleavage gastrulation organogenesis metamorphosis gametogenesis |
|
|
germ layers
|
ectoderm mesoderm endoderm |
|
|
ectoderm turns into what...
|
neuron (CNS) melanocytes (pigment cells) epidermal cells (skin) |
|
|
Mesoderm turn onto what...
|
notochord Bone tissue tubule of kidney RBC facial muscle |
|
|
Endoderm turn into what...
|
stomach cell thyroid cell lung cell |
|
|
Germ cells
|
sperm egg |
|
|
Phylotypic stage
|
vertebrates are visually similar at neurula stage
|
|
|
Epithelial cells
|
cell polarity cell adhesion stationary high levels of E cadherin low levels of N cadherin |
|
|
Mesenchymal cells
|
NO cell polarity loss of cell adhesion migrate/invade low levels of E cadherin high levels of N cadherin |
|
|
Genomic Equivalence
|
All cells of a particular organism contain the same DNA (differential expression of genes within each cell creates morphological and functional diversity)
|
|
|
Enhancer
|
Not required for transcription but increases the speed. directs time, location, amount of transcription. Studied by reporter contruct.
|
|
|
Promoter
|
CRUTIAL for transcription NO promoter = NO transcription DNA adjacent to the gene directs location of transcriptional initiation |
|
|
Histone methylation
|
methylated by: Histone methyltransferases Methyl groups removed by: histone demethylases Activates or represses, depending on other histone modifications Methyltransferases can be recruited by MeCP2 (which interacts with methylcytosine). |
|
|
DNA methylation
|
methylated by: DNMT (DNA de novo Methyl Transferases) Generally, represses transcription Patterns copied to newly replicated DNA and inherited by daughter cells. |
|
|
Mediator
|
30 subunit protein complex that interacts with general transcription factors and specific transcription factors to loop chromatin and initiate transcription
|
|
|
two ways that microRNAs block translation
|
RISC- RNA induced silencing complex- such small regulatory RNAs can bind with the 3’ UTR of messages and inhibit their translation. translational repression |
|
|
Major mechanisms used for cytoplasmic RNA localization.
|
Diffusion and local anchoring Localized protection Active transport along the cytoskeleton |
|
|
in situ hybridization
|
labeled antisense mRNA probe (a DNA or RNA sequence that is complementary to the sequence of a specific mRNA) is hybridized with the mRNA in the organ itself
|
|
|
involved in blocking translation in the amphibian oocyte
|
Maskin
|
|
|
mechanisms used by miRNAs to control mRNA translation
|
Blocking initiation factors Cleaving mRNA Recruiting endonuclease |
|
|
mechanisms of cytoplasmic mRNA localization
|
Active transport along cytoskeleton Localized protection Diffusion and local anchoring |
|
|
Regulation at the transcriptional level
|
Chromatin stateEnhancers/SilencersInsulatorsTranscription factors Mediator Complex & other promoter-enhancer links Promoter typeDNA Methylation status |
|
|
Regulation: Differential RNA Processing
|
RNA Processing: Alternative Splicing
|
|
|
Protein Players (trans-acting factors):
|
U1 snRNA=recognizes 5’ end U2AF=3’ end recognition SF2 (alternative splicing factor) interact with 5’ end, Other SFs interact with branchpoint and other splicing factors |
|
|
RNA sequences (cis-acting sequences):
|
3’ Splice Site (3’SS) 5’ Splice Site (5’SS) Branch Point (BP) |
|
|
Controlling Translation
|
Differential mRNA longevityStored mRNAsRibosomal selectivitymiRNAsCytoplasmic localization
|
|
|
What is the difference between a miRNA and an siRNA?
|
Source of RNA and effects; siRNA is from a non-endogenous source, can be from experimental manipulation (often called RNAi) or from virus while miRNA is encoded in the genome. Effects: siRNA always results in cleavage of the target mRNA, miRNA usually results in translation block especially when the sequences are not a perfect match, when miRNA is a perfect match, it is likely that the mRNA will be cleaved.
|
|
|
Lin-14 and lin-4
|
First identified microRNA and targetControl early developmental timing of c. elegans embryoHomologs in other animalsLin-4 miRNA recognizes several sequences within the UTR of lin-14.
|
|
|
Post-Translational Modifications
|
Cleaved proteinsAssembly into unitsIon bindingCovalent modifications (phosphate, acetyl groups, methyl groups)
|
|
|
E-Cadherin
|
Cadherin protein that expresses on epidermis
|
|
|
Conclusions based on Holtfreter findings
|
Re-aggregated cells became spatially segregated The final positions reflect embryonic positions(selective affinity) Different tissues bind with different affinities |
|
|
property associated with cadherin molecules
|
They are a transmembrane molecule Adhesion strength is related to the quantity of cadherin molecules Not all cadherin sub-types can bind each other |
|
|
Cell adhesion
|
the properties that determine which cells will selectively associate with each other….based on molecules expressed on the cell plasma membrane Differential cell affinity |
|
|
Cell migration
|
the ability of a cell or groups of cells to move through space….accomplished through modification of the cytoskeleton
|
|
|
Cell signaling
|
how cells communicate with each other through chemical and physical factors
|
|
|
Calcium dependent cell adhesion molecules
|
requires extracellular calcium for interaction
|
|
|
E- cadherin
|
present in mammalian embryonic cells and later in epithelial cells
|
|
|
N- cadherin
|
high developing neural cells |
|
|
P- cadherin
|
high in placenta
|
|
|
R- cadherin
|
critical for retina formation
|
|
|
Homophilic binding
|
One cadherin molecule can only bind to a cadherin molecule of it’s own type
|
|
|
cell migration
|
Polarization: Cell senses and recognize extracellular signals, then reorganize its cytoskeleton in order to create a front and a back. Protrusions: Actin polymerization at the cell’s leading edge (the front)FilopodiaLamellipodiaAdhesion of the cell to its extracellular substrate: focal adhesions Release of adhesion in the rear, the cell move in the Forward direction |
|
|
Extracellular Matrix (ECM):
|
made up of fribronectin, collagen, laminin, proteoglycans (heparan and chondroitin suflate)
|
|
|
Fibronectin
|
binds to integrin receptors found in cells
|
|
|
Integrins
|
act as adhesion molecule, linking cytoskeletal elements such as vinculin and talin to microfilaments
|
|
|
permissive induction
|
the responding cell or tissue is already specified (differentiated enough that it only needs extracellular matrix or nutrients as an inducer to continue to its final form).
|
|
|
permissive induction
|
Responder is already specified for a cell fateNeeds only the right environment (eg ECM or nutrients) or signal for expression of previously specified traitsResponding cell’s only ‘choice’ is to develop in the predetermined way or to do nothing.
|
|
|
Instructiveinduction
|
Responding cell fate is not determined yetInducer sends signal that alters gene expression in the responding cellWithout induction, cell cannot differentiate in that particular way (eg removal of optic vesicle in amphibian experiment)
|
|
|
Regional specificity
|
instructive induction Mesenchyme induces epidermal epithelium |
|
|
Genetic Specificity
|
example frog with newt balancers and newt with frog suckers Mesenchyme induces ectoderm to become jaw and external oral structures. |
|
|
RTK pathway
|
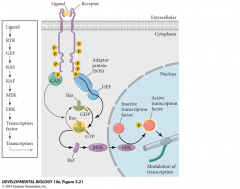
Ligands-FGF, EGF, PDGF, VEGF, etc Receptors-FGFR, PDGFR, VEGFR, etcCrosstalk between ligands and receptors.GEF: GTP exchange factor, activates RasGAP: GTPase activating protein, returns Ras to inactive state |
|
|
JAK-STAT pathway
|
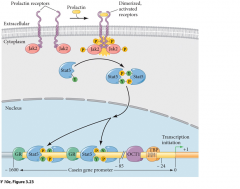
receptors-prolactin, EPO, Thrombopoeitin, leptin, interlukin, GM-CSF promoteligand activates JAK autophosphorilation (activates self) STAT binds and again phosphoilation. STAT dimerizes, enter nucleus, bind to specific regions and promote translation. |
|
|
Hedgehog pathway
|
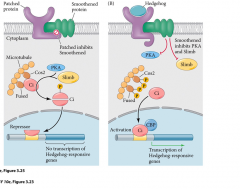
Cyclopamine blocks pathway |
|
|
three types of WNT pathways
|
Canonical-transcriptional controlNon-canonical- cytoskeletal and transcriptional changesCalcium-mediated- transcriptional controlFocus here on Canonical and Non-canonical (sometimes called Planar Cell Polarity)
|
|
|
WNT Pathway
|
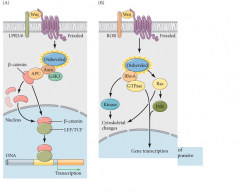
|

