![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
118 Cards in this Set
- Front
- Back
|
What are the ingredients for de novo synthesis of pyrimidine ring?
|
Bicarb-->carbamoyl Pi
Gln (gives the NH3) Asp (C's and N-1 |
|
|
Describe Carbamoyl Phosphaste Synthetase (CPS)
|
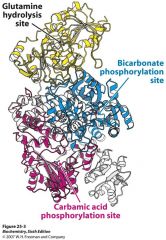
Heterodimeric enzyme with 3 rxns:
Smaller chain Gln hydrolysis for NH4+ Larger chain = 2 ATP grasp domains --Bicarb to carbamic acid in one --Carbamic to carbamoyl phos in other |
|
|
How are the C-N bonds formed in CPS?
|
via acyl-Pi intermediates
Activated by phos from ATP and then the amine group boots Pi and takes its place |
|
|
Where does the NH3 come from in CPS action?
|
hydrolysis of Gln to Glu + NH3
dyad of Cys + His in smaller chain of CPS |
|
|
What enzyme catalyzes the reaction carbamoyl Pi to carbamolyaspartate?
|
Asp transcarbamoylase
|
|
|
What is the reaction catalyzed by pyrimidine phosphoribosyltransferase?
|
Orotate + PRPP-->Orotidylate
|
|
|
What's the enzyme used in the last step to get UMP?
|
Oritidylate decarboxylase
|
|
|
What is CTP formed from?
|
UMP. It must be converted to UTP via kinases (UMP kinase and nucleoside diphos kinase)
|
|
|
How are nucleosides made interconvertible?
|
through phosphorylation by specific kinases (UMP kinase) and broader kinases (nucleoside diphos kinases.
|
|
|
How is CTP made from UTP?
|
carbonyl group is replaced by an amine group (from Gln)
|
|
|
What's the difference between pyrimidines and purines?
|
Pyrimidines (just a circle, less complicated) are made first and then attached to the ribose.
Purines (2 rings) are built onto the ribose as it gets assembled. |
|
|
What are 2 salvage enzymes for making purines from pieces?
|
--Adenine phosphoribosyl transferase (AMP)
--Hypoxanthine-guanine phosphoribosyl transferase |
|
|
What is the salvage enzyme for making uracil + PRPP?
|
pyrimidine phosphoribosyl transferase
|
|
|
What is the difference between the ATP grasp zones of CPS and phosphoribosyl amidotransferase?
|
A Cys residue hydrolyzes the Gln.
phosphoribosyl amidotransferase needs both substrates (Gln and PRPP) in order to work. |
|
|
What does adenosylsuccinate synthase use to activate inosinate?
|
GTP
|
|
|
What does GMP synthetase use to activate inosinate?
|
ATP (adds an AMP, not a Pi to activate)
|
|
|
How are deoxyribonucleotides synthesized?
what is the enzyme? |
reduction of ribonucleotide by NADPH
ribonucleotide reductase |
|
|
What is a nucleoside?
What is a nucleotide? |
purine or pyrimidine base + sugar
Phosphate ester of a nucleoside |
|
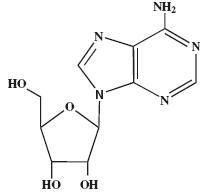
What is this? Is it a nucleoside or nucleotide?
|
adenosine
nuclueoside |
|
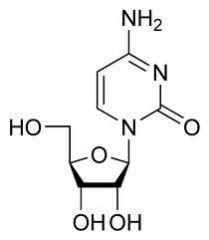
What is this?
Is it a nucleoside or nucleotide? |
cytidine
nucleoside |
|
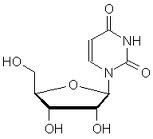
What is this?
Is it a nucleoside or a nucleotide? |
uridine
nucleoside |
|
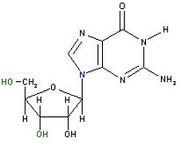
What is this?
Is it a nucleoside or a nucleotide? |
guanosine
nucleoside |
|
|
What is adenylate?
|
nucleotide of adenosine(Phos'd via ester bond)
|
|
|
What is guanylate?
|
nucleotide of guanosine (phos'd via ester bond)
GMP |
|
|
What is Uridylate?
|
nucleotide of uridine
(phos'd via ester bond) UMP |
|
|
What is Cytidylate?
|
nucleotide of cytidine
(phos'd via ester bond) CMP |
|
|
What are the RNA ribonucleosides?
|
adenosine
guanosine uridine cytidine |
|
|
What are the DNA nucleosides?
|
deoxyadenosine
deoxyguanosine Thymidine deoxycytidine |
|
|
What are the RNA nucleotides?
|
adenylate
guanylate uridylate cytidylate |
|
|
What are the DNA nucleotides?
|
deoxyadenylate
deoxyguanylate Thymidylate Deoxycytidylate |
|
|
In pyrimidine, where do the C-2 and N-3 atoms come from?
What about N-1 and C-4.5.6? |
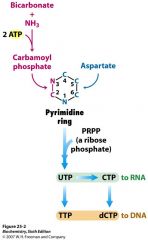
C-1 and N-3 are from carbamoyl Pi and N-3 and C-4.5.6 are from ASP
|
|
|
Where does the NH3+ come from to provide CPS with the N-3 it needs in forming pyrimidine?
|
Gln.
The smaller chain part of CPS has a Cys and a His that hydrolyzes Gln to provide the NH3+ |
|
|
What 3 reactions take place in CPS and where?
What atoms do they provide to the formation of pyrimidine? |
smaller chain=hydrolysis of Gln to give NH3+ to carbamoyl Pi (N-3) of pyrimidine
Larger chain has 2 ATP grasp domains where bicarb is phos'd and other where carbamic acid is phos'd to carbamoly pi provides C-1 |
|
|
What does Asp transcarbamoylase do?
|
puts Asp on carbamoyl Pi
makes carbamoyl Asp |
|
|
Describe what happens once you have carbamoyl Asp.
|
carbamoylAsp --> dihydroorotate + H2O
Dehydration dihydroorotate + NAD+ --> orotate + NADH Oxidation |
|
|
Once you have orotate then what happens?
What is the enzyme that catalyzes the rxn? |
pyrimidine phosporibosyltransferase plops the orotate onto PRPP and makes orotidylate
|
|
|
Once you have orotidylate then what happens?
What is the enzyme that catalyzes this reaction? |
Orotidylate decarboxylase
loose a CO2 from orotidylate and forms UMP (uridylate) |
|
|
Draw the structures and cofactors and enzymes of how carbamoyl Pi gets made into orotate.
|
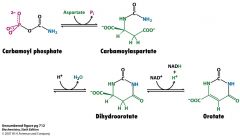
enzyme is Asp transcarbamoylase
|
|
|
draw the structures, cofactors and enzymes of forming orotidylate from orotate in the formation of pyrimidines
|
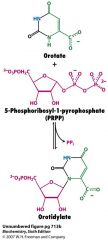
pyrimindine phosphoribosyltransferase (non-specific enzyme) catalyzes the orotate + PRPP part
|
|
|
Draw the structure and cofactors for the decarboxylation of orotidylate to UMP.
What is the enzyme? |
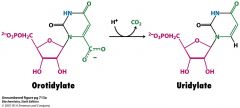
enzyme is orotidylate decarboxylase
|
|
|
What is UMP kinase?
What's the difference between it and nucleoside diphosphate kinase? |
UMP kinase is very specific for making UMP into UDP and creating ADP in the process
Now once you have 2 diphosphates the nucleoside diphosphate kinase comes in and transfers the Pi around (broad specificity) Makes UTP which can then stay as UTP or be made into CTP (animation of UTP) |
|
|
What gets replaced when the amination of UTP happens to make CTP?
What's the mechanism? |
carbonyl group
Gln gives us the NH3 and the O-4 atom of UTP is phos'd to activate it. Then the NH3 is a NU and attacks the activated carbonyl group |
|
|
draw the structures and cofactors and enzymes for the formation of CTP from UTP
|
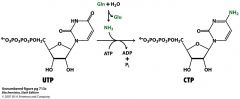
no enzyme for this bad boy
|
|
|
What is the precursor to adenylosuccinate and xanthylate?
|
IMP
|
|
|
What gets released as AMP is being made from Inosinate (IMP)?
|
fumarate
|
|
|
Where does the NH2 come from in AMP?
Which carbon does it go on? |
comes from Asp
goes on C-6 |
|
|
Where does the NH2 come from in GMP?
Which carbon does it go on? |
comes from Gln
Goes on C-2 |
|
|
What is the enzyme that transforms Inosinate into AMP?
|
adenylosuccinate synthase
(utilizes GTP, not ATP) |
|
|
What is the enzyme that transforms IMP to GMP?
|
GMP synthetase
NAD+ is cofactor, then traditional phos and Nu attack by NH3 released from Gln |
|
|
What is the reductant in ribonucleotide reductase?
|
NADPH
|
|
|
Describe the R1 subunit of ribonucleotide reductase.
|
active site
3 Cys and 1 Glu reduce ribose to deoxyribose 2 allosteric control sites |
|
|
Describe the R2 subunit of ribonucleotide reductase.
|
contains the tyrosyl radical which handles the unpaired electron in the aromatic ring
|
|
|
How is thymine made?
|
methylated uracil.
Uses thymidylate synthase |
|
|
What are the 2 purine salvage enzymes?
|
adenine phosphoribosyltransferase
adenine + PRPP-->adenylate + PPi hypoxanthine-guanine phosphoribosyltransferase (HGPRT) guanine + PRPP--> guanylate + PPi hypoxanthine + PRPP--> inosinate (precursor to A &G) + PPi |
|
|
What is the pyrimidine salvage enzyme?
|
pyrimidine phosphoribosyltransferase
uracil + PRPP-->uridylate + PPi |
|
|
Where does N-1 in purine come from?
|
Asp
|
|
|
Where does C-2 in purine come from?
|
N10-formyl THF
|
|
|
Where does N-3 in purine come from?
|
Gln
|
|
|
Where does c-4,5 and N-7 come from in purine?
|
Gly
|
|
|
Where does C-6 in purine come from?
|
CO2
|
|
|
Where does C-8 come from in purine?
|
N10 THF
|
|
|
Where does N-9 in purine come from?
|
ribose P
|
|
|
What are the first and second steps in de novo synthesis of purines?
|
Displacement of PPi from PRPP by NH3 from Gln to create PHOSPHORIBOSYL-AMINE
Glutamine phosphoribosyl amidotransferase is the enzyme Then Gly's carboxyl group is activated via ATP and attached to phosphoribosyl-amine's amine group |
|
|
What is kicked off in purine synthesis when Asp is inserted?
|
fumarate
|
|
|
What's the precursor product at the end of the 9 steps of purine de novo synthesis?
|
inosinate IMP
|
|
|
What is the reactant that when Asp is added, fumarate gets kicked off?
|
5-aminoimidazole-4-(N-succinylcarboxamide) ribonucleotide
|
|
|
What is the base used in salvage synthesis of inosinate?
|
hypoxanthine + PRPP--> inosinate + PPi
Inosinate is the precursor to adenylate and guanylate |
|
|
Describe how AMP is synthesized from Inosinate. Be able to draw structure, know cofactos and enyzme.
|
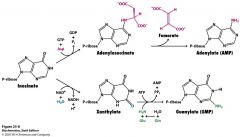
adenylosuccinate synthase is enzyme. Inosinate is already oxidized so the NH from Asp comes in and kicks off fumarate.
|
|
|
How is GMP made from inosinate? Be able to draw structures and know cofacters and enzymes.
|
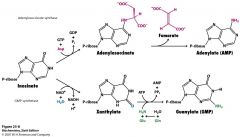
|
|
|
Which Carbon gets the NH3 added to in inosinate by adenylosuccinate synthase?
What is required? Where does the ammonia come from? |
C-6
GTP Asp is where the NH3 comes from |
|
|
Which Carbon does the NH3 get added to in inosinate by GMP synthetase? What is required? Where does the NH3 come from?
|
C-2
NAD+ to oxidize to xanthylate ATP to activate xanthylate NH3 comes from Gln hydrolysis |
|
|
What is the enzyme that works on all ribonucleotides to de-oxyfy them?
|
ribonucleotide reductase
|
|
|
Which C gets deoxy'ed?
What is the reductant (not the enzyme)? |
C-2
NADPH via radical mechanism |
|
|
Where does the radical get generated in order for ribonucleotide reductase to do its thang?
|
nearby Fe iron center
|
|
|
Once the nearby Fe+ center generates a free radical for reducing ribonucleotides, where in the enzyme does it go and on which residue?
|
Gets delocalized on a Tyr ring in the R2 dimer of ribonucleotide reductase
|
|
|
What is the first step in the mechanism of ribonucleotide reductase?
|
transfer of a free radical from a Cys to the tyrosyl ring in R2
|
|
|
What is the enzyme that reduces the disulfide bond formed in the reduction of ribonucleotides?
|
thioredoxin
|
|
|
How is thymine different from uracil?
|
thymine is a methylated uracil
|
|
|
Where does the methyl come from in thymine synthesis?
What is the enzyme that catalyzes this reaction? |
methyl group comes from N5, N10 methylene THF
thymidylate synthase is the enzyme |
|
|
What is the mechanism behind thymidylate synthase?
|
The enzyme attaches a thiol to C-6 which rearranges the electrons and makes c-5 a better Nu to attack N5, N10 methylene THF
Also, thymidylate synthase is also torquing THF to open ring |
|
|
how is dihydrofolate formed?
|
It's the last step in making thymidylate. A transfer of an H to the newly added carbon on dUMP gives you thymidylate and dihydrofolate.
|
|
|
What enzyme regenerates THF after the formation of thymidylate?
|
dihyrofolate reductase uses NADPH to put a H+ back on THF after thymidylate is made.
|
|
|
what is fluorouracil?
|
targets thymidylate synthase in cancer therapy
acts by suicide inhibition because it binds up thymidylate synthase because it acts like a normal substrate initially, but the F+ clogs up the cycle when it cannot be abstracted. |
|
|
how does methotrexate work?
|
analogs of dihydrofolate (amethopterin and aminopterin) bind dihydrofolate reductase and thus THF cannot be regenerated.
|
|
|
What is pyrimidine biosynthesis regulated by?
|
Aspartate transcarbamylase
recall that the key step is bringing carbamoyl phos together with asp to make carbamoylaspartate. |
|
|
How is pyrimidine synthesis inhibited?
How is it activated? |
by the final product initiated by asp transcarbamoylase which is CTP
Activated by large amounts of ATP |
|
|
How is purine biosynthesis inhibited?
|
Gln phosphoribosyl amidotransferase (remember this enzyme kicks off PPi from PRPP and replaces it with an amide from gln just like the name says)
|
|
|
What inhibits Gln phosphoribosylamidotransferase?
|
IMP, GMP and AMP
|
|
|
What activates Gln phosphoribosyl amidotransferase?
|
PRPP activates
|
|
|
What inhibits adenylosuccinate synthase?
|
AMP
|
|
|
What inhibits GMP synthetase?
|
GMP
|
|
|
Describe what reciprocal substrate relation means.
|
Think about inosinate. It can either go to make AMP or GMP. If you notice, adenolysuccinate synthase needs GTP to make AMP and similarly GMP synthetase needs ATP to make GMP so there becomes this balance of utilizing guanine and adenine to make ribonucleotides
|
|
|
Describe what happens if you have a lot dATP with regard to regulation of ribonucleotide reductase and the "overall" allosteric control site.
|
dATP means there is enough reduced ribonucleotides so it inhibits ribonucleotide reductase in the R1 subunit allosteric site.
similarly, if there is a lot of ATP this keeps ribonucleotide reductase in the on switch, we need to make more DNA. |
|
|
What happens when dATP or ATP binds to the substrate specificity allosteric control site of ribonucleotide reductase?
|
changes ribonucleotide reductase to make more CTP and UTP
|
|
|
What inhibits adenylosuccinate synthase?
|
AMP
|
|
|
What inhibits GMP synthetase?
|
GMP
|
|
|
Describe what reciprocal substrate relation means.
|
Think about inosinate. It can either go to make AMP or GMP. If you notice, adenolysuccinate synthase needs GTP to make AMP and similarly GMP synthetase needs ATP to make GMP so there becomes this balance of utilizing guanine and adenine to make ribonucleotides
|
|
|
Describe what happens if you have a lot dATP with regard to regulation of ribonucleotide reductase and the "overall" allosteric control site.
|
dATP means there is enough reduced ribonucleotides so it inhibits ribonucleotide reductase in the R1 subunit allosteric site.
similarly, if there is a lot of ATP this keeps ribonucleotide reductase in the on switch, we need to make more DNA. |
|
|
What inhibits adenylosuccinate synthase?
|
AMP
|
|
|
What inhibits adenylosuccinate synthase?
|
AMP
|
|
|
What inhibits GMP synthetase?
|
GMP
|
|
|
What happens when dATP or ATP binds to the substrate specificity allosteric control site of ribonucleotide reductase?
|
changes ribonucleotide reductase to make more CTP and UTP
|
|
|
What inhibits GMP synthetase?
|
GMP
|
|
|
Describe what reciprocal substrate relation means.
|
Think about inosinate. It can either go to make AMP or GMP. If you notice, adenolysuccinate synthase needs GTP to make AMP and similarly GMP synthetase needs ATP to make GMP so there becomes this balance of utilizing guanine and adenine to make ribonucleotides
|
|
|
Describe what reciprocal substrate relation means.
|
Think about inosinate. It can either go to make AMP or GMP. If you notice, adenolysuccinate synthase needs GTP to make AMP and similarly GMP synthetase needs ATP to make GMP so there becomes this balance of utilizing guanine and adenine to make ribonucleotides
|
|
|
Describe what happens if you have a lot dATP with regard to regulation of ribonucleotide reductase and the "overall" allosteric control site.
|
dATP means there is enough reduced ribonucleotides so it inhibits ribonucleotide reductase in the R1 subunit allosteric site.
similarly, if there is a lot of ATP this keeps ribonucleotide reductase in the on switch, we need to make more DNA. |
|
|
Describe what happens if you have a lot dATP with regard to regulation of ribonucleotide reductase and the "overall" allosteric control site.
|
dATP means there is enough reduced ribonucleotides so it inhibits ribonucleotide reductase in the R1 subunit allosteric site.
similarly, if there is a lot of ATP this keeps ribonucleotide reductase in the on switch, we need to make more DNA. |
|
|
What happens when dATP or ATP binds to the substrate specificity allosteric control site of ribonucleotide reductase?
|
changes ribonucleotide reductase to make more CTP and UTP
|
|
|
What happens when dATP or ATP binds to the substrate specificity allosteric control site of ribonucleotide reductase?
|
changes ribonucleotide reductase to make more CTP and UTP
|
|
|
How does the binding of TTP impact ribonucleotide reductase?
|
More GDP gets reduced and inhibits pyrimidine synthesis
|
|
|
How does GTP impact ribonucleotide reductase?
|
ATP will get reduced to dATP
|
|
|
How are nucleotides degraded to nucleosides?
And further to Ribose 1-Pi and free bases? |
nucleotidases
nucleoside phosphorylases |
|
|
What does phosphoribomutase do?
|
makes ribose 1-Pi to ribose 5-Pi
Remember ribose 5-Pi is a substrate for making PRPP |
|
|
Name 3 enzymes that degrade AMP.
|
nucleotidases
adenosine deaminase nucleoside phosphorylase |
|
|
What is SCID? What physiological effects does it have?
|
Severe combined immunodeficiancy
low levels of adenosine deaminase recurring infections lead to death without gene therapy loss of T cells Too much dATP and ribonucleotide reductase is inhibited so no DNA synthesis |
|
|
What does xanthine oxidase do ?
Why is this significant theraputically? |
oxidizes hypoxanthine to xanthine and ultimately urate
allopurinol is an analogue for hypoxanthine and thus binds up xanthine oxidase so less urate is made |
|
|
Why would high levels of urate in primates be advantageous from a genetic standpoint?
|
urate acts as a ROS scavenger--good antioxidant.
|
|
|
What is Lesch-Nyhan Syndrome?
|
deficiency in hypoxanthine guanine phosphoribosyltransferase which is a salvage enzyme.
increase in PRPP, purine biosyn, urate compulsive, self-destructive aggressive behavior in children and kidney stones and gout later |

