![]()
![]()
![]()
Use LEFT and RIGHT arrow keys to navigate between flashcards;
Use UP and DOWN arrow keys to flip the card;
H to show hint;
A reads text to speech;
55 Cards in this Set
- Front
- Back
|
Bacteria often respond to environmental change by regulating |
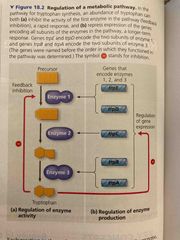
Transcription A cell can regulate the production of enzymes by feedback inhibition or gene regulation |
|
|
Operon model |
Mechanism for control of gene expression A cluster of functionally related genes controlled by a single on/off switch |
|
|
Operator |
Segment of DNA usually position within the promoter Acts as a on/off switch |
|
|
Operon |
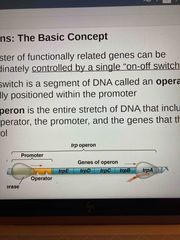

Entire stretch of DNA, includes the operator, promoter, and genes they control |
|
|
Repressor |
A protein that can switch off the operon Prevents gene transcription by binding the operator thus blocking RNA polymerase Product of regulatory gene located distance away from operon |
|
|
Regulatory gene |
Encodes a repressor protein Has its own promotor and is some distance away from operon Expressed continuously at low rates |
|
|
A repressor can be in an active or inactive form depending on... |
Presence of other molecules A corepressor is a molecule that works with a repressor protein to switch operon off ie: E.coli can synthesize tryptophan when it has insufficient tryptophan |
|
|
By default, the trp operon is... |
On and genes for tryptophan synthesis are transcribed When tryptophan is present, it binds to the trp repressor protein, turning off operon |
|
|
The repressor is active only in the presence of its... |
Corepressor. Thus the trp operon is turned off of tryptophan levels are high |
|
|
Repressible operon |
One that is usually on Binding of repressor to the operator shuts off transcription ie: trp operon |
|
|
Inducible operon |
One that is usually off An inducer (molecule) inactivates the repressor and turns on transcription ie: lac operon |
|
|
Lac operon |
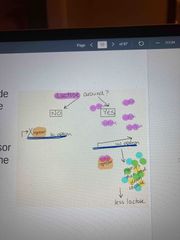
Inducible operon Contains genes that code for enzymes used in hydrolysis and metabolism of lactose |
|
|
By itself, the lac repressor is ____ and switches the lac operon _____ |
Active and off |
|
|
Inducer |
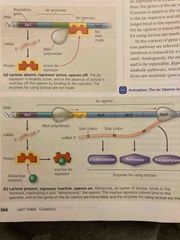
A molecule that inactivates the repressor to turn the lac operon on |
|
|
Inducible enzymes usually function in ____pathways |
Catabolic Their synthesis is induced by a chemical symbol |
|
|
Repressible enzymes usually function in a _____pathway |
Anabolic Their synthesis is repressed by high levels of the end product |
|
|
Negative control |
Gene Regulation of both trp and lac Operon are switched off by the active form of their repressor protein |
|
|
Positive control |
Gene regulation involving a regulatory protein that acts directly with the genome to switch transcription on |
|
|
Positive control gene regulation |
Through a stimulatory protein (cyclic AMp receptor protein) When glucose is scarce, CRP is activated by binding with cAMP |
|
|
Cyclic AMP receptor protein(CRP) |
An activator of transcription Positive control of gene regulation |
|
|
CRP process |
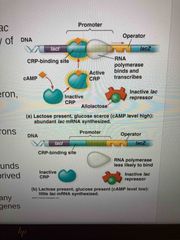
Low glucose, CRP is activated Attached to promotor of lac operon and increases affinity of RNA polymerase (more transcription) When glucose levels increase, CRP detaches from lac operon and transcription returns to normal rate |
|
|
CRP helps regulate other operons that encode enzymes used in... |
Catabolic pathways |
|
|
The ability to catalyze compounds like_____enables cells, deprived of glucose, to survive |
Lactose Compounds present in any cell determine which genes are switched on |
|
|
Genes are turned on/off in response to signals from their... |
External and internal environments All organisms must regulate what genes are expressed at any given time In multicellular, gene expression regulation is essential for cell specialization (each cell must maintain a specific program of gene expression) |
|
|
Differential gene expression |
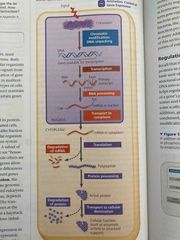
The expression of different genes by cells with the same genome Abnormalities in gene expression can lead to cancer Regulated at many stages but often equated with transcription |
|
|
A human cell expresses about ___Of its protein-coding genes at a given time |
20% Highly differentiated cells (muscle or nerve) express smaller fractions of their genes |
|
|
Structural organization of chromatin helps regulate gene expression in several ways |
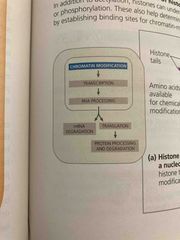
Genes with highly packed heterochromatin are usually not expressed Chemical modifications to histones and DNA of chromatin influence both chromatin structure and gene expression |
|
|
Histone modifications and DNA methylation |
Histone acetylation Methylation DNA methylation Can be passed on to future generations of cells |
|
|
Histone acetylation |
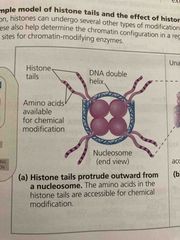
Acetyl groups are attached to an amino acid in a histone tail Add acetyl group makes more hydrophilic This opens up chromatin structure, promoting initiation of transcription |
|
|
DNA methylation |
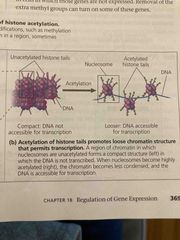
Addition of methyl groups to certain DNA bases (histone tails) Become more hydrophobic Associated with reduced transcription in some species (condenses chromatin reducing transcription) Can cause long term inactivation of genes in cellular differentiation |
|
|
In genomic imprinting, methylation regulates expression of |
Maternal or paternal alleles of certain genes |
|
|
Epigenetic inheritance |
The inheritance of traits transmitted by mechanisms not directly involving then nucleotide sequence “On top of the gene” |
|
|
Regulation of transcription initiation |
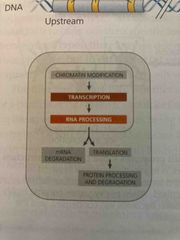
Chromatin-modifying enzymes provide initial control of gene expression They make a region of DNA more or less able to bind the transcription machine |
|
|
Control elements |
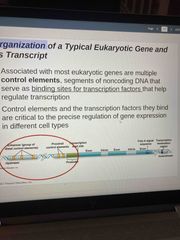

Segments of noncoding DnA in most eukaryotic genes binding sites for transcription factors Help regulate transcription |
|
|
Control elements |
Segments of noncoding DnA in most eukaryotic genes binding sites for transcription factors Help regulate transcription |
|
|
Critical to the precise regulation of gene expression in different cell types |
Control elements and their transcription factors that bind |
|
|
General transcription factors |
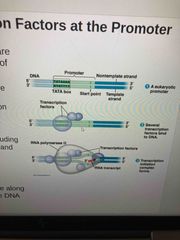
Essential for transcription of all protein-coding genes ie: RNA polymerase needs it to initiate transcription, some bind to TATA box in promoter |
|
|
General transcription factors |
Essential for transcription of all protein-coding genes ie: RNA polymerase needs it to initiate transcription, some bind to TATA box in promoter |
|
|
Only when the complete____ has assembled can RNA polymerase begin to move along the template strand of the DNA |
Initiation complex |
|
|
Close to the promoter |
Proximal control elements |
|
|
Enhancers |
Distant control elements Far away from a gene or even located on an intron Each enhancer is associated with only one gene |
|
|
An activator is a protein that binds to____and stimulates transcription of a gene |
An enhancer Have 2 domains: one that binds to DNA and a second that activates transcription |
|
|
Protein-mediated bending |
Current model suggest DNA is bent to bring bounds activators into contact with a group of mediator proteins This helps assemble and position the preintiaition complex |
|
|
Combinatorial control of gene activation |
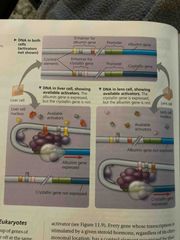
A particular combo of control elements can activate transcription Only when the right activator proteins are present With only a dozen or so control elements, a large number of combos are possible |
|
|
Coordinately controlled genes in eukaryotes |
Co expressed eukaryotic genes are not organized in operons (some exceptions) Can be scattered over different chromosomes but each has same combo of control elements Activator proteins in the nucleus recognize specific control elements and promote simultaneous transcription of the genes |
|
|
Chromatin from different chromosomes... |
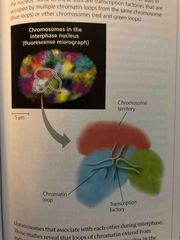
Congregate at particular sites (rich in transcription factors and RNA polymerase) Chromosomes conformation capture techniques allow id of regions of chromosomes that interact with each other |
|
|
Regulatory mechanisms can operate after transcription? |
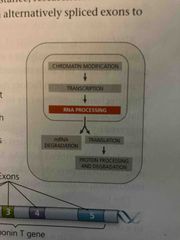
Yes, transcription alone does not consistute gene expression Fine-tune gene expression rapidly in response to environmental changes |
|
|
Alternative RNA splicing |
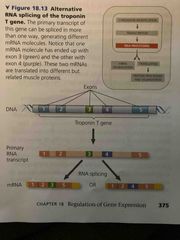
Different mRNA molecules are produced from same primary transcript pending which segments are Exons or introns More than 90% of the human proteins-coding genes undergo alternative splicing |
|
|
Expands significantly the repertoire of a eukaryotic genome |
RNA splicing It is a proposed explanation for the surprisingly low number of genes in the human genome |
|
|
Blocked intimation of translation for selected mRNAs |
Regulatory proteins, miRNA that binds sequences, structures of mRNA Alternatively, translation of all mRNAs in a cell may be regulated simultaneously (ie: translation initiation factors are simultaneously activated in an egg following fertilization |
|
|
____mRNa is more long-lived than ______mRNa |
Eukaryotic than prokaryotic Life span of mRNA molecules in the cytoplasm is important in determining the pattern of protein synthesis in a cell |
|
|
Resides in the untranslated region (UTR) at the 3’ end of the molecule |
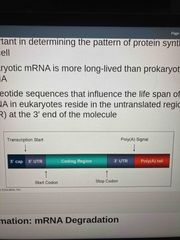

Nucleotide sequence that influences life span of mRNA in eukaryotes |
|
|
After translation, polypeptides undergo processing |
Protein processing and degradation Cleaves and chemical modifications |
|
|
Length of time each protein functions is regulated by |
Selective degradation Cells mar proteins by attaching ubiquitin to them This is recognized by proteasomes which degrade the proteins |
|
|
In a prokaryotic cell, when there is low glucose, there is high... |
Cyclic AMP levels CRP detects cAMP levels and then activate lac operon |

